|
Split a multi-molecule file into new1.smi, new2.smi, etc.: xnįor further specific information and options, use -HĬonversion from a SMI file in STDIN to a Mol2 file written to STDOUT: Output format options are preceded by 'x', e.g. Input format options are preceded by 'a', e.g. Individual file formats may have additional formatting options. The following formats are currently supported by Open Babel:Īrc - Accelrys/MSI Biosym/Insight II CAR format Ĭache - CAChe MolStruct format Ĭacint - Cacao Internal format Ĭar - Accelrys/MSI Biosym/Insight II CAR format Ĭdx - ChemDraw binary format Ĭom - Gaussian Cartesian Input Ĭrk2d - Chemical Resource Kit 2D diagram formatĬsr - Accelrys/MSI Quanta CSR format įch - Gaussian formatted checkpoint file format įchk - Gaussian formatted checkpoint file format įck - Gaussian formatted checkpoint file format įh - Fenske-Hall Z-Matrix format See -H format-ID for options allowed by a particular format -v SMARTSĬonvert only molecules NOT matching SMARTS pattern specified -VĬompress the output with gzip File Formats Separate disconnected fragments into individual molecular records -tĪll input files describe a single molecule -title titleĪdd or replace molecular title -x optionsįormat-specific output options. pĪdd Hydrogens appropriate for pH (use transforms in phmodel.txt) -propertyĪdd or replace a property (e.g., in an MDL SD file) -s SMARTSĬonvert only molecules matching the SMARTS pattern specified -separate Specifies output format, see below for the available formats -O outfile Splitting one input file - put each molecule into consecutively numbered output filesīatch conversion - convert each of multiple input files into a specified output formatįor multiple entry input, stop import with molecule # as the last entry -o format-ID Translate computational chemistry modeling keywords (e.g., GAMESS and Gaussian) -m Join all input molecules into a single output molecule entry -k Specifies input format, see below for the available formats -j -join Output formatting information and options for all formats -i Output formatting information and options for format specified -Hall Output the available fingerprint types -h 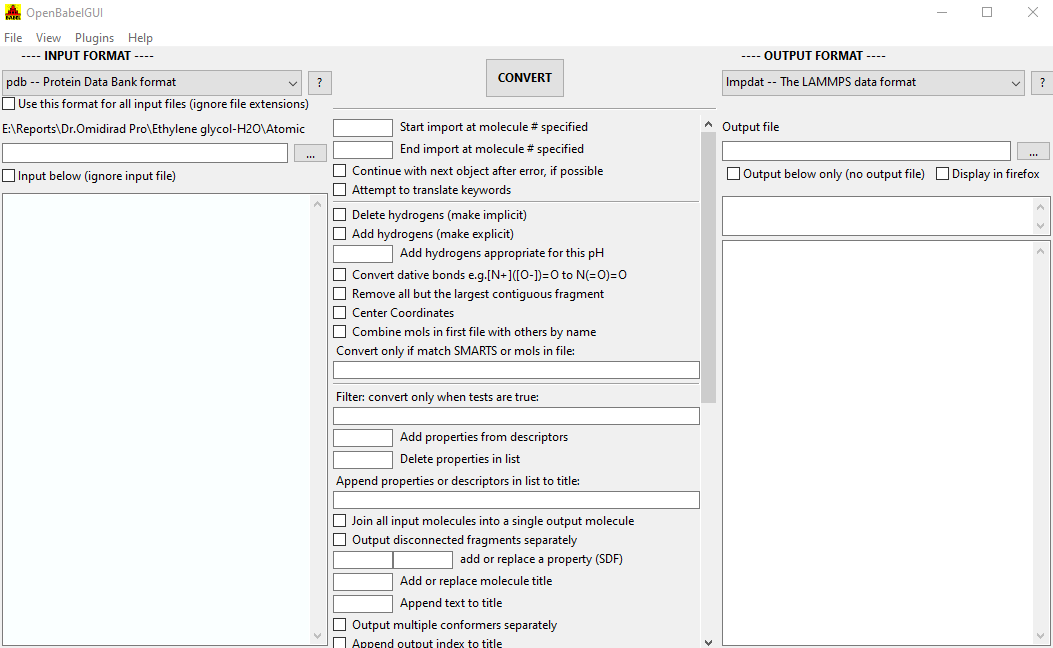
See -H format-ID for options allowed by a particular format -addtotitleĪppend text to the current molecule title -addformulaĪppend the molecular formula after the current molecule title -bĬonvert dative bonds: e.g., ()=O to N(=O)=O -cĬombine molecules in first file with others having the same name -eįilter the level of errors and warnings displayed:Ĥ = include "audit log" messages of changes to dataįor multiple entry input, start import with molecule # as the first entry -F More than one can be used, and a molecule title can be included if enclosed in quotes. 
The SMILES string should be enclosed in quotation marks. 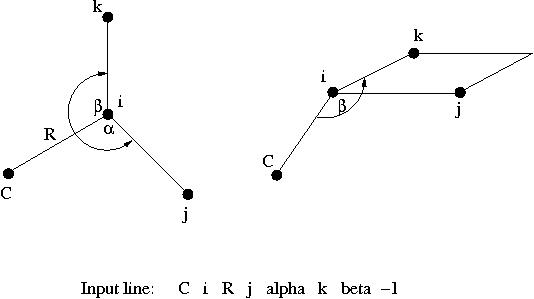
: " SMILES-string"Įnter SMILES string and use it in place of an input file. If only input and output files are given, Open Babel will guess the file type from the filename extension. 
For more information, se the Open Babel web pages. Open Babel is also a complete programmers toolkit for developing chemistry software. Obabel is a cross-platform program designed to interconvert between many file formats used in molecular modeling and computational chemistry and related areas.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
